Why Brain Emulation is Coming Sooner Than Many Think – A BPF Response to Dr. Miller’s NYT Editorial on Brain Emulation
Dr. Kenneth D. Miller, Co-Director of the Center for Theoretical Neuroscience at Columbia University, in a recent editorial in the New York Times, “Will You Ever Be Able to Upload Your Brain?”, recently raised several skeptical arguments about the possibility of brain uploading. A key point to take away from this essay is that Dr. Miller actually agrees that successful brain emulation is possible and consistent with our current understanding of the brain. He says:
“I am a theoretical neuroscientist. I study models of brain circuits, precisely the sort of models that would be needed to try to reconstruct or emulate a functioning brain from a detailed knowledge of its structure. I don’t in principle see any reason that what I’ve described could not someday, in the very far future, be achieved.”
While Miller acknowledges the theoretical possibility of brain emulation, he is very skeptical about any near-term ability to do so, claiming that successfully emulating a human brain is not only infeasible with current and impending technologies (a claim no one would dispute), but that it will also vastly outstrip technological capabilities far into the future. We at the Brain Preservation Foundation (BPF) certainly agree that brain emulation presents tremendous challenges. For example, Kenneth Hayworth, President of the BPF, has described the resources and effort needed to emulate a human brain as akin to a “moon shot project,” meaning that society would have to devote significant resources to the project over decades. However, such a projection contrasts starkly against Miller’s suggestion that such a project could take “thousands or even millions of years”.
Two very promising techniques for brain preservation have recently been developed by BPF prize contenders, one by Dr. Robert McIntyre and another by Dr. Shawn Mikula. While it is still an open question, these methods are likely to preserve the connectome and all the molecular detail necessary for a person’s memory and identity; this is an exciting area of current research. Far from being pessimistic, we have every reason to be hopeful that validated and reliable human brain preservation protocols will be developed in the next decade. Memory and identity theory and emulation research, for its part, will also continue to advance, and we have many reasons to expect validated models for long-term memory storage, beginning in well-studied circuits in simple model organisms, again within the next decade.
One reason for our considerably more optimistic predictions is that brain emulation relies on computing and other information technologies, fields that have classically enjoyed exponential gains in capability and should continue to do so for the foreseeable future. Computational neuroscience is now big science, and significant resources, above and beyond previous investments and commitments, are now being devoted to emulation. See for example the the EC-funded Human Brain Project. In basic neuroscience, new tools and funding have emerged for mapping and understanding the connectome and the “synaptome,” Dr. Stephen Smith’s term for the deep synaptic diversity that exists at each synaptic bouton. See his presentation, The Synaptome Meets the Connectome, 2012 (below), to appreciate how great this diversity is, and how incomplete our models of it are to date. Fortunately, new funding for synaptic characterization, including the NIH-funded BRAIN Initiative, has also emerged. This increase in research and funding is driven by the need to better understand complicated mental illnesses as well as the general goal of bettering our understanding of the brain in both its structural and functional aspects. Therefore, our knowledge of the workings of the neuron and brain will continue to improve.
Miller also raises doubts as to whether it will ever be possible to preserve the brain for later scanning and emulation and states “It will almost certainly be a very long time before we can hope to preserve a brain in sufficient detail” for accurate brain emulation. Miller suggests that successfully emulating the brain may require extraordinary detailed molecular knowledge of the state of every synapse and dendrite. It is an ongoing question just how much detail is required to emulate a neuron, and it is not a foregone conclusion that dynamics at the scale of individual molecules are necessary; it is entirely possible that such molecular-scale properties are simply not critical to the larger stochastic behaviors of neurons. But if molecular-level detail for scanning and emulation turns out to be necessary, a number of promising new protocols, such as 21st Century Medicine’s Aldehyde-Stabilized Cryopreservation, may be sufficient to the task, as aldehyde-stabilization via cerebrovascular perfusion is a process that can rapidly lock down all molecular activity in neurons within minutes after death. This and other emerging protocols may preserve cellular ultrastructure throughout an entire mammalian brain, a claim the BPF was founded to assess. Accurate whole brain preservation is the central focus of the BPF, and rather than being a distant possibility it may soon be a reality.
Returning to emulation, another reason to be optimistic that it is feasible sooner than many expect is that from an engineering perspective, the brain is a very noisy system, and therefore unlikely to rely on details at the scale of individual molecules, and it relies on considerable redundancy of structure and function. In contrast to what Miller suggests, cutting edge models of neurons currently used by computational neuroscience groups such as the Blue Brain Project (for the paper, see Markram etal. Cell 2015 Oct 8;163(2):456-492) model hundreds of aspects of the neuron using the software NEURON (PDF here) and not “a single fixed strength” for synapses. Miller further claims the “wash of chemicals from brainstem neurons that determine such things as when we are awake or attentive and when we are asleep, and by hormones from the body that help drive our motivations” are problematic, but it is not clear why improved models of the brain can’t include such properties in the future, if they are necessary. Miller further claims:
“dendrites and synapses (and more), are constantly adapting to their electrical and chemical ‘experience,’ as part of learning, to maintain the ability to give appropriately different responses to different inputs, and to keep the brain stable and prevent seizures”
“The connectome might give an average strength for each connection, but the actual strength varies over time.”
Again, we see no reason why all of these aspects, if relevant, would not be able to be accounted for in future large-scale models of the brain if they turn out to be necessary.
Furthermore, when we do neural emulation of any kind, we must also ask, “emulation for what purpose?” What features will we need to capture and “upload” from neurons to preserve their highest-level information? What subset of all neural structure and processes encodes, for example, our life’s episodic memories? Neuroscience is still only beginning to ask such questions, which are critical to evaluating the future of uploading.
For example, Henry Markram’s Blue Brain Project (BBP), mentioned above, is one leading computational neuroscience effort that seeks to emulate neural electrophysiology. It is largest slice of rat somatosensory cortex that has been emulated to date. In a recent minireview of the BBP (see, A Biological Imitation Game, Cell 2015 Oct 8;163(2):277-80), neuroscientists Christof Koch and Michael Buice of the Allen Instititute for Brain Science note that this emulation is an “impressive initial scaffold that will facilitate asking [specific and quantifiable] questions of brains.” The BBP may allow us to ask the deepest questions yet about what data are necessary to emulate cortex electrophysiology, and what data can be excluded. The BBP currently uses deterministic Hodgkin-Huxley partial differential equations as its “lowest level of granularity” to represent neural firing activity. Koch and Buice ask whether, for example, stochastic Markov models of “thousands of tiny [ion] channel conductances” might also be needed to model the subtleties of neural activity and coordination. The BBP simulation does not at present model ion channels, perhaps due to the current limitations of the IBM Blue Gene supercomputer they use for the emulation. It is important to note that the Markram team presently thinks that emulating ion channel detail will be unnecessary to reproduce neural electrophysiology. Koch and Buice point out that the only way to know if such detail is needed will be to conduct a Turing Test-like “imitation game.” If the emulation gives the same response as wet neurophysiological experiments, at the level of system performance needed by the electrophysiologist, then the emulation has succeeded. Presumably such tests will be forthcoming from Markram’s lab and others in coming years.
What is not yet clear in these emulation efforts, perhaps because we don’t yet know enough about neural information storage to even ask this question well, is what level of detail will be needed not only for electrophysiology emulation, but also for high-level memory encoding and retrieval. Fortunately, an “imitation game” for some of this kind of information is also being played today not only by neuroscientists and computational neuroscientists, but also by computer scientists, including those working with biologically-inspired architectures like deep learning neural nets. Computer scientists are racing to figure out how to store high-level information like episodic memories in biologically-inspired associational networks on their own, computational neuroscientists are racing to emulate all neural activity, and neuroscientists are racing to “imitate,” via better static and dynamic descriptions, all the remaining still-poorly-characterized features of neural structure and activity, such as ephaptic coupling and perineuronal nets.
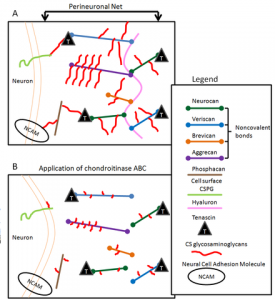
Perineuronal Nets – Extracellular matrix structures responsible for synaptic stabilization in the adult brain (Wikipedia 2015).
Each of these races will inform each other, and at some point, we will know the subset of stable features and processes arising from neural anatomy (stable to all the chaos and trauma that affects live brains) that encode the high-level information we care about most. Perineuronal nets, for example, may regulate synaptic stability and our learning ability, but they may turn out not to be part of our stored learned knowledge itself. They may be one of those features of neural activity that we would like, but don’t “need” to emulate, for brain preservation to be a personally and socially valuable activity. As our knowledge advances, we will need to ask some important questions much better than we have been doing to date.
We will need to ask questions like: What kind of information in our own lives do we care most about preserving? If we were forced to choose, some of us might start with episodic memories, and move down from there to the hundreds of distinct “neural modules” of cognition, emotion, and personality. Stated another way, if preservation services were affordable, accessible, and validated, and a few of your friends had already done it at the end of their own lives, how much of “you” would you need to reasonably expect would be preserved in order to make the brain preservation choice? If you only had a reasonable expectation that your higher memories would be preserved, and nothing else, would that be enough? If you expected you would lose your perineuronal nets, due to (let us presume, for the sake of argument) damage to the extracellular matrix in the preservation process, and so expected be revived with all your unique memories, but the loss of your unique learning proclivities (being instead given “species average” perineuronal nets on your revival) would that be enough? If you were brought back like Henry Molaison after his hippocampal lesion, so that you had only your memories up to your death, and had to be given a hippocampal replacement (a “species average” module for your short-term memory), would that be enough? If you lost your epigenomic data, and that turned out to take away, let’s say, 10% of the personality features that made you different from your identical twin sibling (the other 90% of your differences being due to your unique memories and neural connections, which were successfully preserved), would that be enough? If you could keep your cortical memories, but your emotional state would be reverted to an earlier “you” on revival, would that be enough? Finally, and most importantly for some, how affordable and reliable can we make the various preservation options, and can they be done sustainably with respect to the environment?
We believe brain preservation is a personal choice each of us should be able to freely make for ourselves, or not, in light of ever-changing science, technology, and self-understanding, at the point of our own deaths. Clearly, brain emulation will never be a perfect process. At best, it will attempt to statistically approximate perfection, as neuroscience and computer science get better at knowing what features of the most complex and fascinating material structures on Earth, our brains, can be preserved, what features are “worth” preserving and reviving, and what features are not. The preservation option already exists today. Seeking to validate that option, or finding that it cannot be validated, is our institutional mission. The more we know, the more informed our choice can be.